
In fact, a survey of lossy quality score compressors has already shown that changing quality scores alone can sometimes have a beneficial effect on variant calling 9. 
Current variant calling procedures are complex and rely on alignment information, including quality scores which are a direct function of the analog signals used to determine the called base. 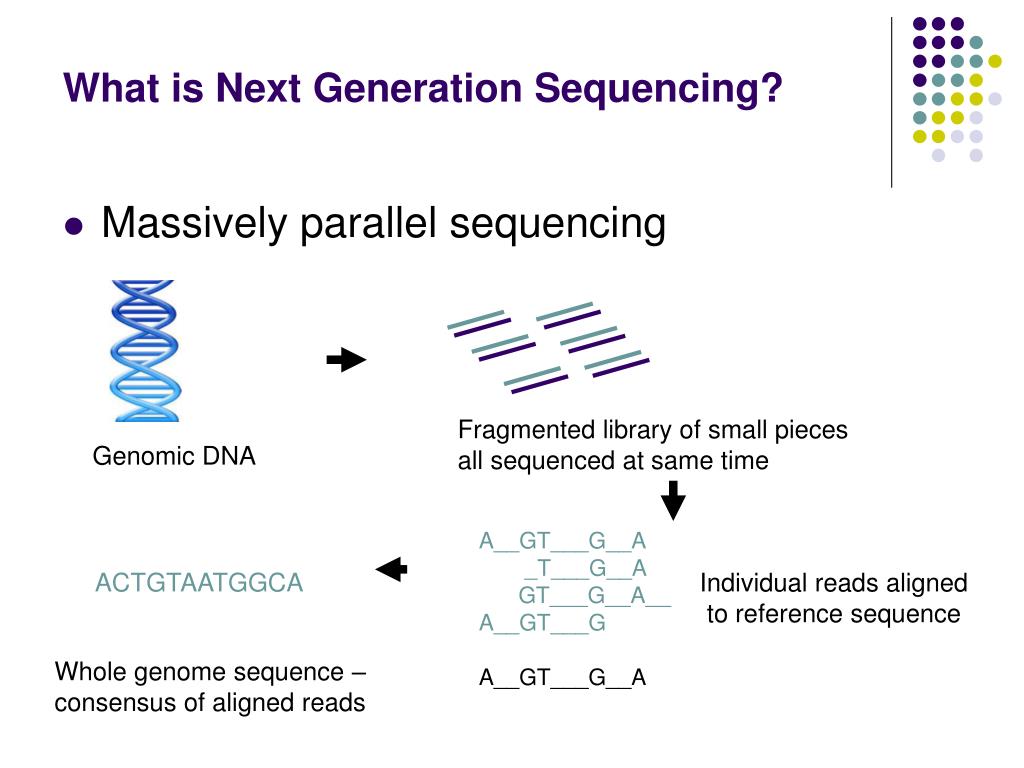
To our knowledge, none of these works examines the effect that denoising might have on variant calling. However, these analyses often do not extend to later steps in genomic sequence analysis pipelines, and those that do focus on non-human data sets 8. They are typically tested on simulated and real data sets in FASTQ format, and have been shown to perform well on some of the early stages of genomic sequencing pipelines, such as correcting base calling errors in the simulated data sets, increasing both breadth and depth of reads coverage during alignment 6, or improving de novo assembly of real data sets 7. These denoisers attempt to rectify sequencing errors by only changing individual bases in reads, while retaining the original quality scores. Note that there is a clear distinction between the natural variations of DNA sequences (i.e., genetic polymorphisms)-which are the target of variant calling-and noise due to sequencing errors introduced by the sequencing platform, which can be represented as base call mismatches (single base-substitutions) and INDELs.Īlgorithms for removing noise, or denoisers, have been proposed for genomic sequencing data 4, as well as for other biological methods relying on genomic sequencing, like ChiP-seq 5. 
Thus, accuracy of variant identification is paramount. Variant identification from WGS is increasingly being used for diagnosis and treatment design in the clinical setting, especially in the field of rare genetic disease research 3. These errors can affect downstream applications, with an important application being variant calling, or the identification of genetic polymorphisms unique to individuals. Furthermore, these errors were found to be correlated with position within the read, resulting in position-dependent noise characteristics. For example, Illumina sequencing technologies produce “short” reads on the order of hundreds of bases, with an average substitution error rate of less than 1%, and INDEL rates orders of magnitude lower 2. However, the genomic sequencing process is imperfect and can result in reads containing various types of noise including base substitutions, insertions, and deletions (INDELs).Īlthough noise characteristics vary across sequencing technologies, they are well characterized for some sequencing platforms. Both file types comprise sequences of nucleotide bases called “reads,” which are accompanied by sequences of quality scores that indicate the sequencing machine’s confidence in the base calls making up the reads. Raw sequencing data are typically stored in the FASTQ file format and converted to the SAM file format following alignment to a reference genome. The ability to sequence genetic material has expanded our understanding of genes and their roles in biological processes, opened up new areas of biological inquiry, and are guiding the trajectory of modern biomedical research 1.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
